Principal Component Analysis (PCA) Explained
Learn how Principal Component Analysis (PCA) reduces the dimensionality of datasets while preserving important information. Understand the intuition, mathematics, and practical uses of PCA in machine learning and data science.
🧊 Principal Component Analysis (PCA)
PCA finds the most important directions in data and compresses the data into fewer numbers while trying to keep the important information
PCA is a dimensionality reduction algorithm that:
- finds directions of maximum variance
- projects data onto lower-dimensional space
- minimizes projection error
Run PCA only on the inputs to learn a mapping:
where:
Original training set:
Becomes:
Now the learning algorithm trains on lower-dimensional data.
Important:
- PCA map data of dimensions into dimensions
- PCA is an unsupervised algorithm
- PCA does not use labels
- Do NOT fit PCA on:
- cross-validation set
- test set
The most widely used algorithm for dimensionality reduction.
Example:
Given a messy training data with
- 1000 features
PCA says:
“Maybe only 20 directions are really important.”
Advantages:
- Speed Up Learning: Lower-dimensional data makes training faster.
- Compression : Reduce storage and memory requirements.
- Visualization: visualization becomes easier for lower dimension
Bad Use of PCA
PCA is NOT a good method for preventing overfitting.
Some people think:
fewer dimensions = less overfitting
but this is not ideal.
Reason:
- PCA ignores labels
- PCA may throw away useful predictive information
Instead:
✅ Use regularization to reduce overfitting.
Do NOT automatically add PCA to every ML pipeline.
Bad habit:
Training Data
↓
PCA
↓
Logistic Regression
↓
Predictions
Before using PCA, first try:
Training Data
↓
Learning Algorithm
↓
Predictions
Use PCA only if:
- training is too slow
- memory usage is too large
- dimensionality is extremely high
How to select in PCA?
A common way to choose the number of PCA components is by checking how much variance is retained.
PCA tries to minimize the projection error:
where:
- = original data point
- = projected/reconstructed point
The total variation in data is:
A standard rule is to choose the smallest such that:
This means:
- projection error
- equivalently, 99% variance retained
People usually describe PCA quality as:
- 99% variance retained
- 95% variance retained
- 90% variance retained
How to select Projection Direction?
A good projection line is one where:
- When we project each point onto the line
- The distance between the original point and its projection is small
⚠️ Projection errors
The orthogonal distance from a point to the line.
- Help with selecting projection direction
The projection error is:
So goal of PCA is to find the direction that minimizes the total squared orthogonal distance.
where:
- is the projected version of
PCA minimizes:
Important:
- If PCA returns or , it does not matter.
- Both define the same line.
General Case: nD → kD
Now suppose:
Where is original number of dimensions
Example for 3D
and we want to reduce to dimensions, eg. when we want to project 3D to 2D
Instead of finding one vector, we find vectors:
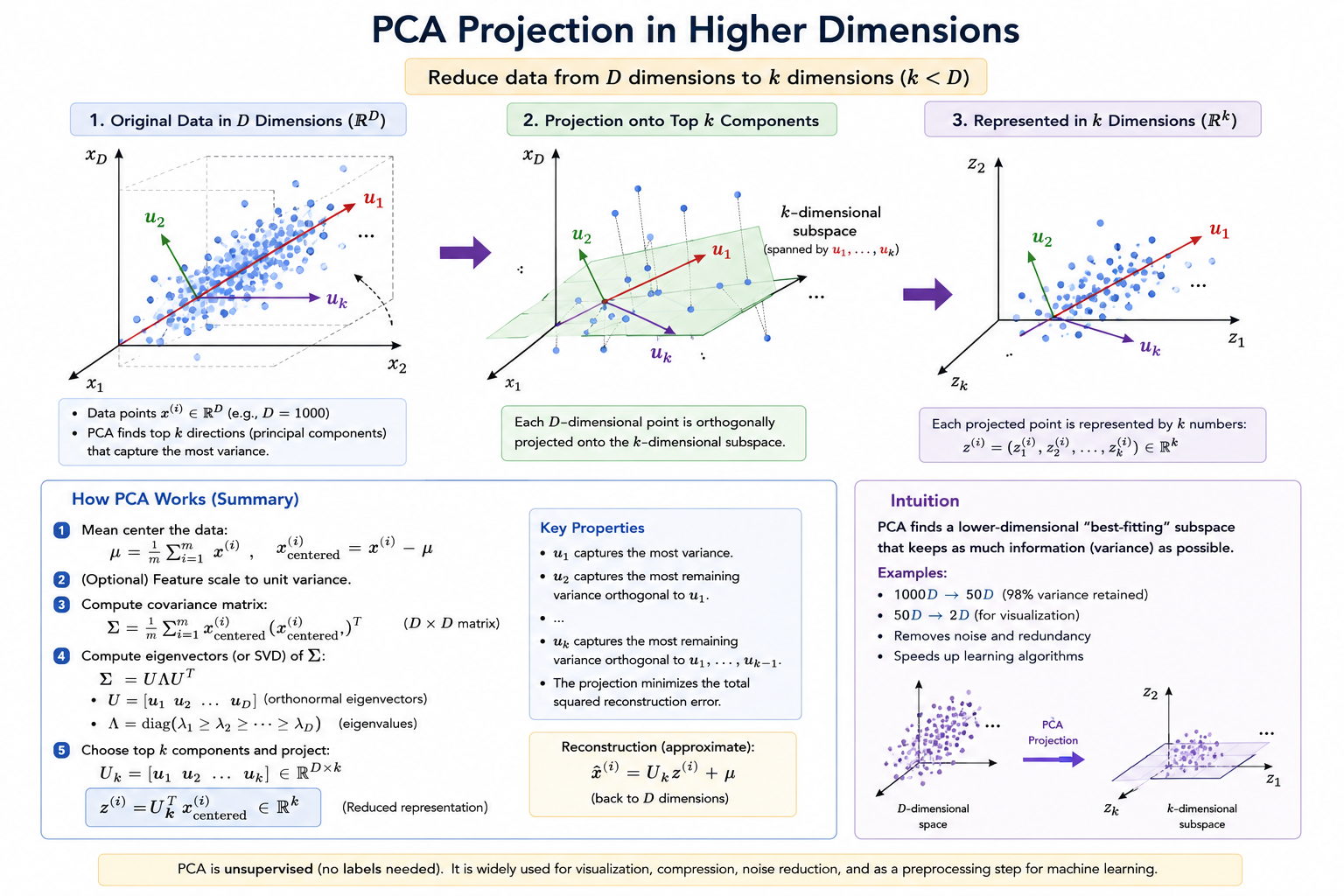
These vectors:
- Define a k-dimensional surface
- Span a k-dimensional linear subspace
We then project each point onto that subspace.
3D → 2D Example
If:
and we reduce to 2D:
- We find two vectors:
- These define a plane.
- Each point is projected onto that plane.
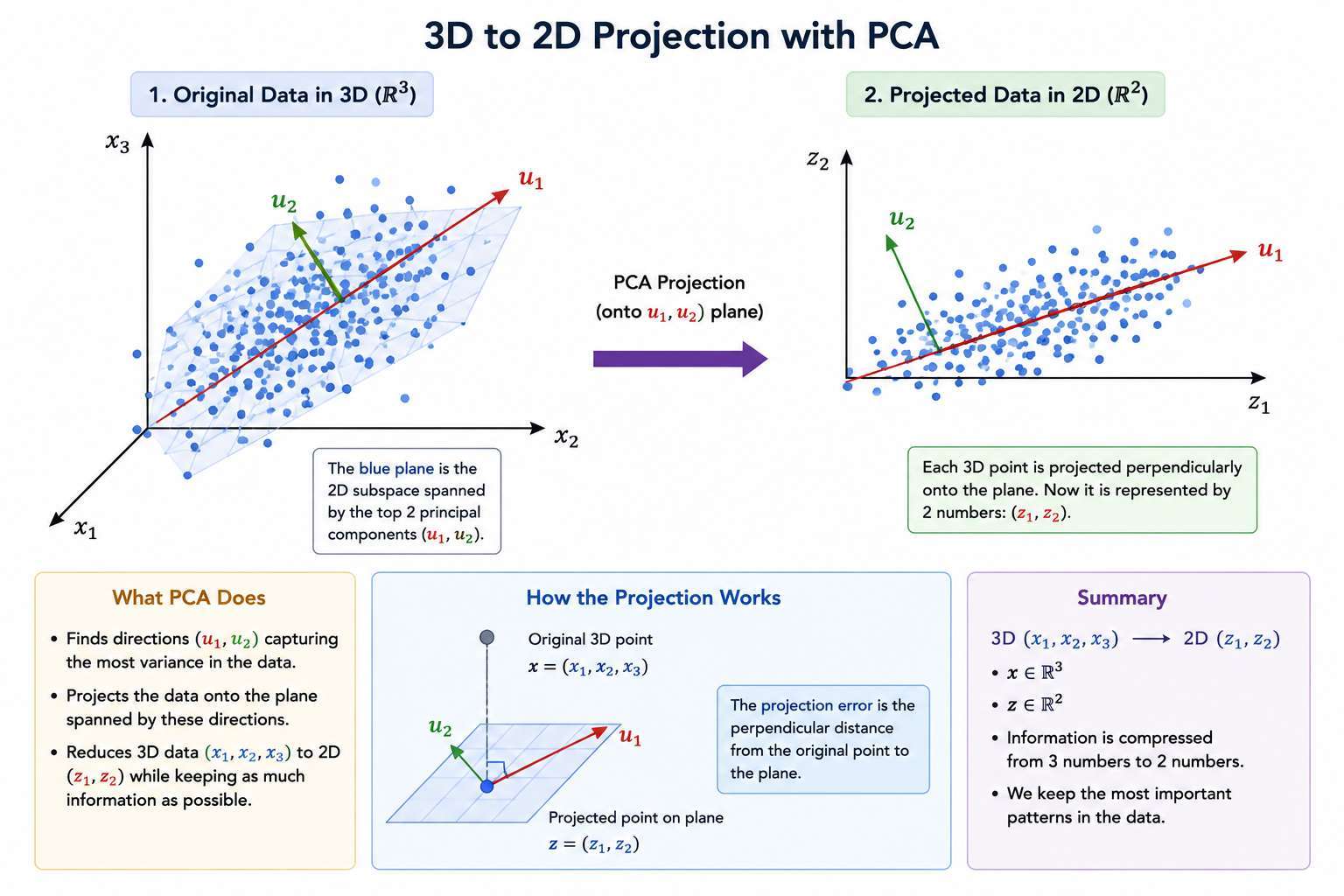
2D → 1D Example
Suppose we have:
and we want to reduce the data from 2 dimensions to 1 dimension.
That means:
- We want to find a line
- Onto which we project all data points
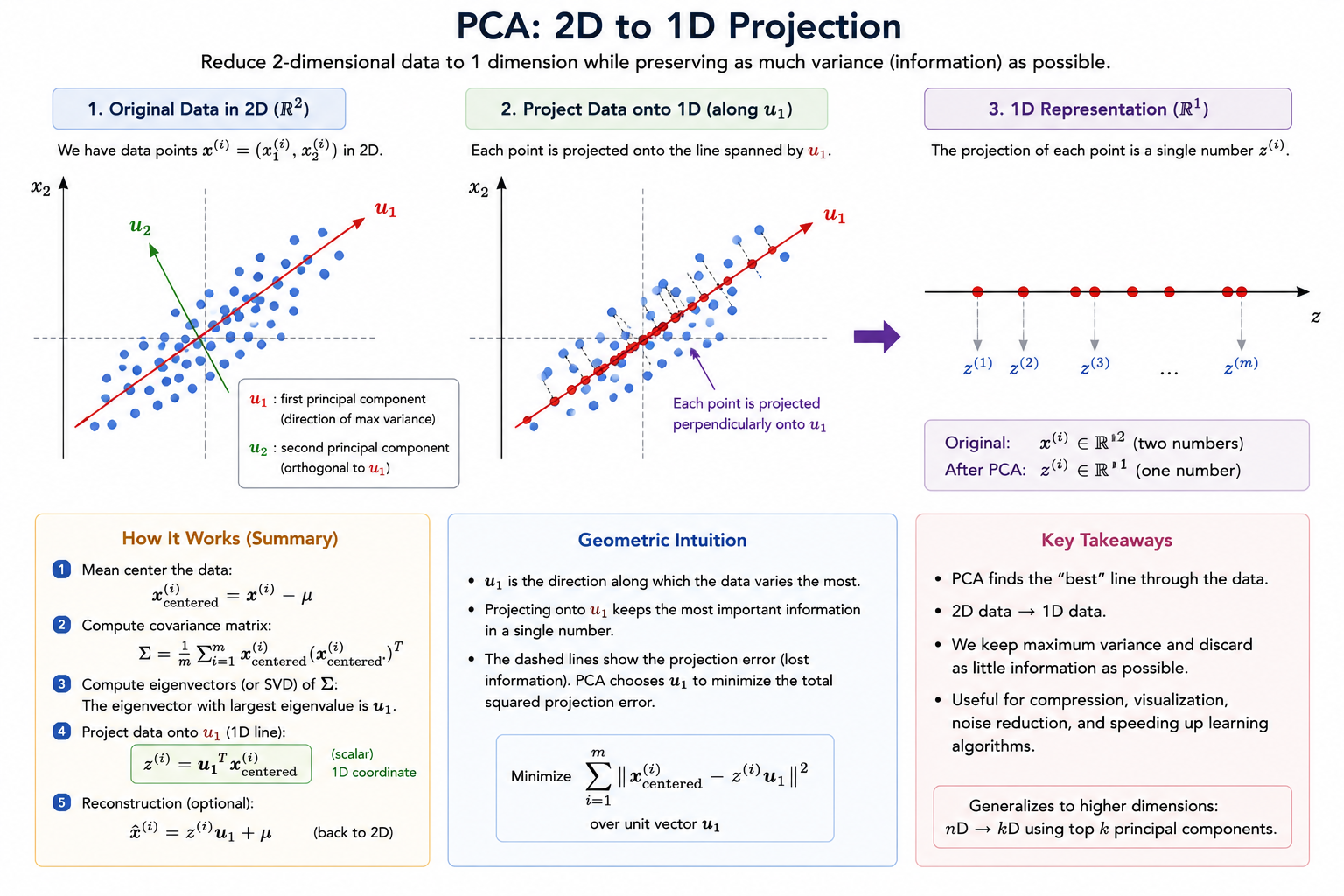
💡 PCA Algorithm
Suppose we have supervised learning data:
where:
- = input features
- = labels
1. Ignore Labels Temporarily
Extract only the input vectors:
Before applying PCA, it is standard to:
1. Perform mean normalization
For each feature:
This makes each feature have zero mean.
2. Perform feature scaling (recommended)
Especially when features have different ranges.
where:
- = mean of feature
- = standard deviation or feature range
So that:
- Each feature has zero mean
- Features have comparable ranges
This prevents one feature from dominating purely due to scale.
Step 2: Compute Covariance Matrix
Covariance Matrix ()
is a square matrix giving the covariance between each pair of elements of a given random vector.
If:
- = number of examples
then covariance matrix is:
Vectorized implementation:
import numpy as np
# Assuming data is (observations, features)
data = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
# Set rowvar=False to treat columns as variables
cov_matrix = np.cov(data, rowvar=False)
print(cov_matrix)
Step 3: Apply Singular Value Decomposition (SVD)
Compute Eugen Vector for Covariance Matrix ()
where
- is dimension matrix.
to take dimensions select first k columns
- = diagonal matrix of singular values
Then variance retained can be computed efficiently as:
Choose the smallest such that:
for 99% variance retained.
Typical values:
- 90%
- 95%
- 99%
Most commonly:
-
95% to 99% variance retained.
-
= right singular vectors (not used in PCA)
import numpy as np
# Define your matrix
A = np.array([[1, 2], [3, 4], [5, 6]])
# Perform SVD
U, S, Vt = np.linalg.svd(A)
print("U (Left Singular Vectors):\n", U)
print("\nS (Singular Values as 1D array):\n", S)
print("\nVt (Right Singular Vectors - Transposed):\n", Vt)
Step 4: Choose Top K Components
Take the first columns:
This reduces data from:
- dimensions to
- dimensions
Step 5: Project Data
Compute reduced representation:
Which is equivalent to
where:
- : represents the original input values in n dimension
- : represents the coordinates of in the reduced k-dimensional space.
Reproduce from given
We know
we can calculate
Where
- is matrix
- is matrix
PCA vs Linear Regression (Very Important)
PCA is NOT linear regression.
Linear Regression:
- Predicts a special variable
- Minimizes vertical squared errors
- Error is measured in the y-direction only
PCA:
- Has no special target variable
- All features are treated equally
- Minimizes orthogonal (shortest) distance to a line/plane
Linear regression minimizes:
PCA minimizes:
These are completely different objectives.
Final Summary
PCA:
- Finds a lower-dimensional subspace
- Projects data onto that subspace
- Minimizes squared orthogonal projection error
- Treats all features symmetrically
- Is not a predictive model
Formally, PCA solves:
where is the projection of onto a k-dimensional subspace.
